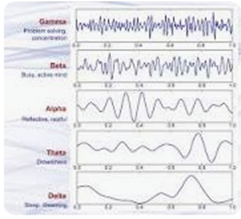
A major challenge for brain researchers is interpreting the results of electroencephalogram (EEG) graphs used to visualize brain activity of everything from meditation to neurological disorders.
Although machine learning can be used, EEG data is extremely multidimensional and can be expensive and time-consuming to annotate, requiring the deep expertise of neurologists and sleep experts. There are insufficient labeled examples for supervised deep neural networks to learn from. A new paper published in the Journal of Neural Engineering demonstrates a promising approach to training deep neural networks to directly learn what EEG looks like, without using labeled data
Although labeled EEGs identifying sleep stages and brain activity are scarce, there is ample unlabeled data. In the paper, Uncovering the Structure of Clinical EEG Signals with Self-supervised Learning, Interaxon researcher Hubert Banville and researchers at Université Paris-Saclay, University of Helsinki, and Max Planck Institute, applied self-supervised learning to extract features from unlabeled EEGs. When limited numbers of labeled data were available, the self-supervised learning approach outperformed traditional supervised learning methods that rely purely on labeled data. The results were obtained on two different EEG classification problems: identifying sleep stages in overnight recordings and detecting EEG pathologies. The approach also uncovered a fascinating structure in the data that relates to clinical information such as sleep stages, pathology, and age.
The research approaches will allow the scientific community, including researchers at Interaxon with one of the largest brain data (EEG) collections in the world, to leverage, mine, and utilize large amounts of unlabeled EEG data to discover relevant information in EEG. This has the potential to improve the performance of algorithms used in everything from consumer sleep and wellness support tools like Muse S, to swifter diagnosis of neurological disorders.